Prediction with Bayesian networks
Introduction
Once we have learned a Bayesian network from data, built it from expert opinion, or a combination of both, we can use that network to perform prediction, diagnostics, anomaly detection, decision automation (decision graphs), automatically extract insight, and many other tasks.
In this article we focus on prediction.
Prediction is the process of calculating a probability distribution over one or more variables whose values we would like to know, given information (evidence) we have about some other variables.
A few examples of predictions are given below. We will also cover a number of advanced prediction concepts and analytical approaches that use prediction as a subroutine.
(t denotes time, C1 & C2 are continuous variables, D1 & D2 are discrete variables)
- P(D1), P(C1)
- P(D1, D2)
- P(C1, C2)
- P(C1, C2, D1, D2)
- P(C1 | D2)
- P(C1, C2[t=0], C2[t=5], D2, D2[t=7])
Inputs and outputs
The variables we are predicting are known as Output variables, while the variables whose information we are using to make the predictions are known as Input variables.
In statistics, Input variables are often called predictor, explanatory, or independent variables, while Output variables are often called Response or dependent variables.
In fact Bayesian networks are more general that dealing strictly with Inputs and Outputs. This is because any variable in the graph can be an input or output or even both. We could even predict the joint probability of an Output and a missing Input. Nevertheless describing variables in terms of their roles as Inputs or Outputs remains a useful concept.
When a model is built from data, predicting outputs is known as Supervised learning.
Missing data
A key feature of prediction, and indeed any activity with Bayesian networks, is that any of the Inputs, whether they are discrete or continuous, can be missing when we perform predictions.
This is useful for a number of reasons:
- We may not currently know the value of an input (e.g. a sensor is broken, and not transmitting, or a patient did not answer a question)
- An anomaly analysis has determined that the value from a defective sensor is clearly wrong, and we therefore want to exclude that value
Note that Bayesian networks natively handle missing due to their probabilistic foundations.
Typically there is no need to fill in missing data via imputation or other methods, although of course
Bayesian networks themselves can be used to fill in missing data.
:::
Multiple outputs
Bayesian networks are not limited to predicting a single output. They can simultaneously predict multiple outputs.
Outputs can be discrete, continuous or a mixture of both :::
Joint prediction
Crucially, Bayesian networks can also be used to predict the joint probability over multiple outputs (discrete and or continuous).
This is useful when it is not enough to predict two variables separately, whether using separate models or even when they are in the same model.
For example, if a network contained continuous outputs C1 & C2, and discrete outputs D1 & D2, we could calculate any or all of the following:
- P(C1, C2)
- P(C1, D1)
- P(C1, C2, D1, D2)
- P(C1, D1 | C2, D2)
- etc...
Distributions
When we make a prediction using a Bayesian network we get a distribution back.
Discrete variables
For a discrete variable D with 3 states {Low, Medium, High} a prediction will be of the form [0.1, 0.3, 0.6]. I.e. the probability of membership of each state.
The task of predicting a discrete variable is often referred to as Classification in line with other approaches.
Continuous variables
For a continuous variable C a prediction will contain both a Mean and a Variance.
The task of predicting a continuous variable is often referred to as Regression in line with other approaches.
Multiple variables
For a joint prediction over multiple continuous variables, the distribution returned will be a multi-variate Gaussian, which consists of a vector of means and a covariance matrix.
For a joint prediction over multiple discrete variable, the distribution returned will be a table containing a probability for each discrete combination.
For a joint containing both discrete and continuous, the distribution returned will be a mixture of multi-variate Gaussians.
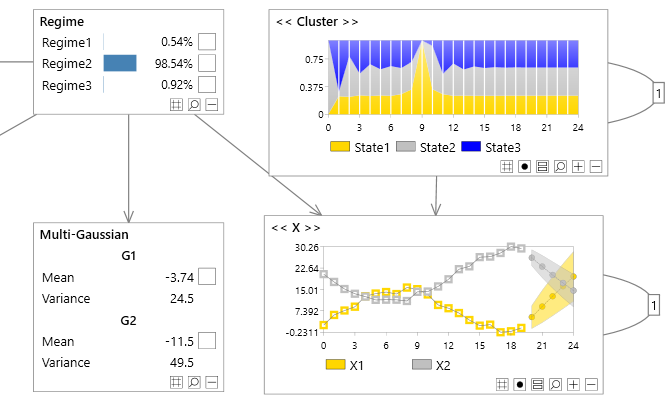
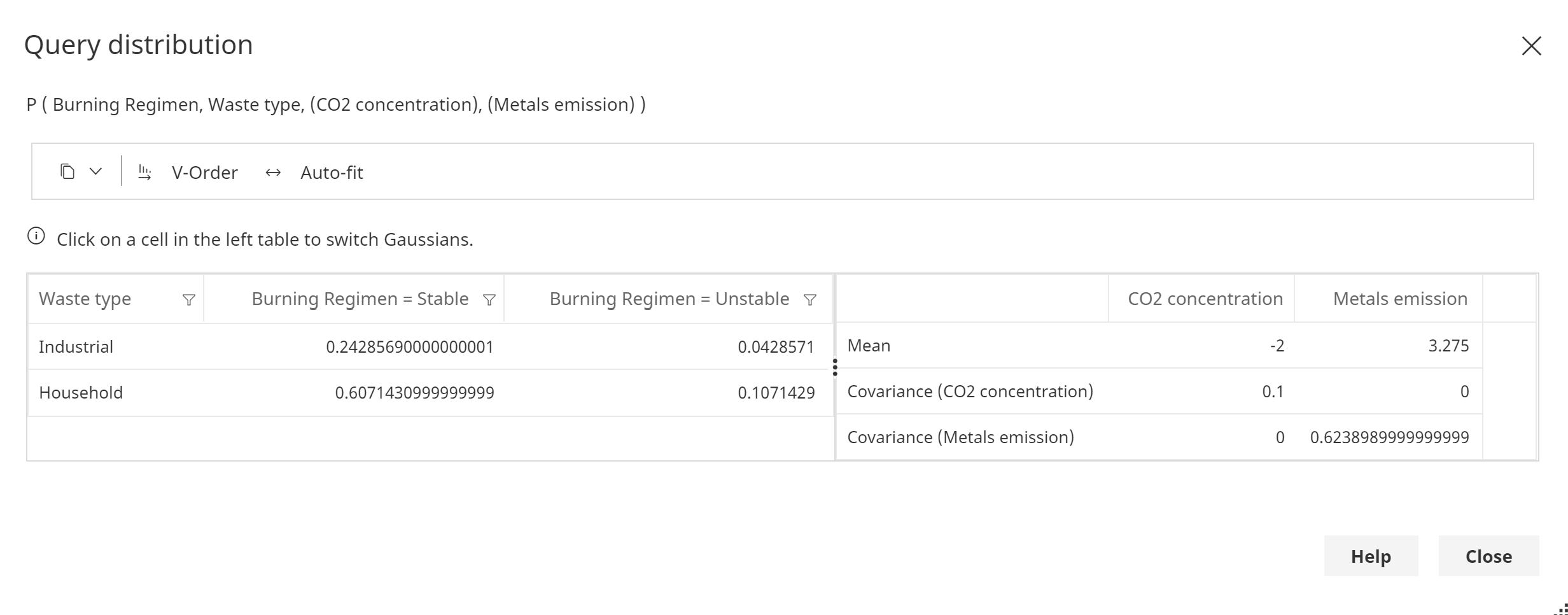 A joint prediction over 2 discrete and 2 continuous variables.
A joint prediction over 2 discrete and 2 continuous variables.
Model evaluation - data
When a Bayesian network has been built from data, it is common place (as with other approaches) to evaluate the performance of the model on unseen data.
There are a number of ways we can do this:
Training and test
Split our data into a training set and a test set. Train a model on the training set and evaluate it on the unseen data in the test set.
Training, test & validation
If we are trying out different options during learning, training and test sets may not be enough, so we can further make use of a validation set, to avoid over-fitting on the test set.
Cross validation
Cross validation splits our data into a number of partitions. For each partition p a model is evaluated on p having been trained on all the data excluding p.
Hence we evaluate our Bayesian network on all of data, but it still remains unseen to each model.
When splitting data, it is common practice to use a random number generator to decide which records (cases) go into which set. However care should be taken with time series or sequence data (data which is not I.I.D) as the data may be dependent in time. A better approach in this case is to ensure one or more contiguous blocks of time series data are used within each partition, and those can be chosen randomly.
Model evaluation - metrics
In order to assess the performance of a Bayesian network, in terms of prediction, we can use the same techniques that are found in other approaches. These are typically split into classification metrics for evaluating the prediction performance of a discrete variable, and regression metrics for continuous.
Classification metrics
Accuracy
Accuracy is simply the percentage of predictions the model got correct.
Another interesting technique is Virtual accuracy.
Confusion matrix
A Confusion Matrix, tells us how well the model performed for each state of the discrete variable.
With binary classifiers (predicting a discrete variable with 2 states, such as True & False), there are names given to particular cells, such as:
- Recall rate
- Sensitivity
- Specificity
- False positive rate
- False negative rate
- True positive rate
- True negative rate
In many scenarios, the False negative rate or False positive rate is more important than the overall accuracy.
Lift chart
Lift charts are another useful tool for analyzing the performance of a classifier.
For a particular state of interest (e.g. Purchase=True), they tell us how the Bayesian network model compares to one which makes perfect predictions and one which make naive predictions. The naive predictions (often called random) are based on the distribution of the variable being predicted.
Lift charts are particularly useful when the state of interest is rare, such as a visitor to a website purchasing an expensive product (Purchase=True). A model can achieve a very good accuracy (e.g. 99%) just by getting the Buy=False cases correct. However we are really interested in how well the model performs over and above a naive model when it predicts Purchase=True.
Regression metrics
For evaluating the performance of a Bayesian network on a continuous variable (regression) we can use a number of regression metrics which are common to other approaches also.
R squared
R-squared, also known as the Coefficient of determination is a standard metric which tells us how well the Inputs explain the variance of the Output.
An R-squared is typically between 0 and 1 (the closer to 1 the better), however can legitimately be negative on unseen data although this indicates a very poor model.
Mean absolute error
The Mean absolute error often shortened to MAE, tells us the average error between the actual value of the output and our predicted value.
Unlike the root mean squared error, the MAE is less sensitive to outliers, so is useful when a small percentage of poor predictions are not important.
Root mean squared error
The Root mean squared error often shortened to RMSE, is similar to the MAE, except it uses squared error values.
Unlike the MAE, RMSE is more sensitive to outliers, therefore is useful when a small percentage of poor predictions are important.
Residuals
When building a regression model, it can be important to look at the residuals.
Residuals are simply the difference between the actual value and the predicted value. They are different to the error, as the sign (positive or negative) is kept.
By plotting residuals, we can see if the residuals look random. If not, it can be an indication of a poor model, as we are missing some important structure.
Temporal residuals
Whether using a time series model or not, when modeling time series or sequence data (non I.I.D. data) it is important to look at how the residuals vary over time (temporal residuals).
If there is structure in the temporal residuals, again it can indicate a poor model, which is not capturing important structure (e.g. seasonality).
Model verification
When making predictions with a Bayesian network (or any other approach), it is important to know whether the new data we are making predictions on is consistent with our model. If our model is built on historic data that no longer reflects reality, should we trust the predictions?
In order to verify that our Inputs are consistent with our model, we can use Anomaly detection.
Before we make a prediction, we can calculate an anomaly score based on our Inputs. If this score looks normal we can continue to make the prediction. If it looks abnormal then:
- We should not trust the prediction
- We could use a missing value in place of the prediction, if being consumed by another process
- We could retrain our model incorporating more recent data to see if that improves matters
Most probable explanation
Instead of standard prediction, it can be useful to predict the most likely configuration of the remaining variables in a Bayesian network that do not have evidence set.
This is known as the Most probable explanation, or for discrete time series variables it is known as the Most probable sequence (a more general version of the Viterbi algorithm).
Retracted evidence
Retracted evidence allows us to see what the prediction on a variable X would have been had X not had evidence set, without affecting any other predictions which continue to use the evidence set on X.
This can be a useful way of monitoring whether our evidence is consistent.
Time series prediction
Bayes Server supports time series and sequence models. In fact we can have models with:
- Multiple interacting continuous time series
- Multiple interacting discrete time series (sequences)
- A mixture of both discrete and continuous time series
- A mixture of both time series variables and standard non time series variables
Models with time series variables are called Dynamic Bayesian networks (DBN).
With a DBN we can make the following types of query:
- Prediction - calculating queries a number of time steps into the future.
- Filtering - calculating queries at the current time.
- Smoothing - calculating queries a number of time steps into the past (calculating historic values)
- Most probable sequence - calculating the most likely sequence of past values (generalized version of the Viterbi algorithm)
- Joint prediction over multiple time series and non time-series variables
The same time series variable A can be included in a joint prediction at different times. e.g. P(A[t=0], A[t=5], B, C[t=1])
Decision automation
We can include one or more utility variables (to model costs or profits for example) and one or more decision variables in a Bayesian network. These models are called Decision graphs.
We can also use Evidence Optimization to perform Decision Automation.
Decision graphs can be used to automate a decision making process. Crucially they make these decisions in an uncertain environment. Uncertain, because we typically do not know the exact state of the system, but just have probabilistic beliefs in various variables.
Decision graphs can include multiple decisions and multiple utilities.
Derived analytics
Predictions on their own are very powerful, but there are also a number of techniques which make use of predictions providing advanced analytics capabilities.
Model structures
In order to make predictions with a Bayesian network, we need to build a model.
A model can be learned from data, built manually or a mixture of both.
Bayesian networks are graph structures (Directed acyclic graphs, or DAGS). There is therefore no fixed structure of a network required to make predictions. Any network can make predictions.
When using data it can sometimes be useful to learn the structure of the network automatically from the data, which is known as structural learning. However we often build the structure manually, taking advantage of well known model types, their extensions, and making use of latent variables.
Latent variables
Latent variables also known as hidden variables, are not directly mapped to data during training. They allow the Bayesian network to automatically capture hidden patterns in the data which have not been directly recorded.
Using one or more latent variables in a Bayesian network (discrete or continuous) often increases the predictive power of the network.
Latent variables are a form of automatic feature extraction and are similar to hidden layers in a Neural network or Deep learning.