Noisy nodes
Introduction
Each node in a Bayesian network requires a distribution which is conditioned on its parents.
For example, if a node C has 3 parents P1, P2 and P3, the distribution required for C is P(C|P1,P2,P3), which can
be read 'the probability of C given P1, P2 and P3'.
When the nodes involved are discrete, the number of parameters required for the child node increase exponentially with the number of parents. For example, consider binary nodes (nodes with 2 states). A binary node with 10 binary parents would require a distribution with 1024 parameters.
In the network below, a node has 100 parents, yet inference can still be performed efficiently using noisy nodes.
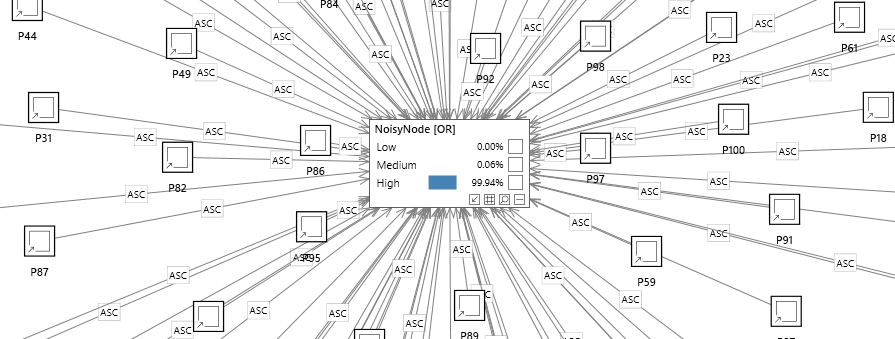
Without noisy nodes, when the number of parents increases it can be difficult to justify each of these parameters. If you were specifying the parameters manually by asking experts, they would need to be able to estimate the probability of the child given all the parents. For example if the child node is a symptom which has 3 parent diseases, the expert would need to be able to answer questions such as 'What is the chance of a patient experiencing a 'fever' given that they have a flu and they have an infection but do not have rheumatoid arthritis. As the number of parents (diseases) increases this becomes more and more difficult and unreliable. The same problem exists when learning the parameters from data. In order to justify each parameter in the model, there needs to be sufficient data 'coverage' for each combination. What's more, inference can become inefficient, perhaps prohibitively so.
There are a number of modeling techniques that can be used to avoid this unwanted explosion of parameters.
- Consider reversing some or all of the links. This is the approach used in a Naive Bayes model, but it can only be used when the conditional independence assumptions can be justified, or when they are a good enough approximation.
- Consider using a technique such as 'divorcing' which recursively groups related parent nodes in order to reduce the number of parameters.
- Consider noisy nodes (causal independence) when you want to model the effect on the child node from each parent node independently, and have their influences combined using logic (e.g. OR logic). The term noisy reflects the fact that the OR combination is probabilistic rather than deterministic, i.e. parents do not have to be on or off, and their effects are combined probabilistically.
Features
Bayes Server supports:
- The representation of noisy nodes via multiple distributions, including a leak distribution.
- Efficient inference with noisy nodes. (We do not expand to the equivalent full distribution, as illustrated in the network shown with 100 parents)
- Efficient parameter learning with noisy nodes.
- Exact and approximate inference with noisy nodes.
- Noisy nodes and parents with 2 or more states (e.g. Noisy Or, Noisy Max).
Creating noisy nodes
A noisy node can be specified in Bayes Server when creating a node, by changing the Noisy type in the New node window. Alternatively the Noisy type of a node can be changed later from the Property editor
Noisy states
The states of a noisy node in Bayes Server should be ordered as follows: { None, Low, ... , High }, for example:
- Text - None, Low, Medium, High
- Text - Absent, Present
- Boolean - False, True
- Numeric - 0, 1, 3, 10
Parameterization
Once a noisy node has been created, a number of distributions need to be specified for it. One for each parent and a leak distribution. The distribution for each parent defines the effect that parent has on the child, whilst the leak distribution allows you to model all other effects.
When parents are linked to a noisy node, the link between them has a Noisy order which determines the order in which parent states affect the child states.
Consider the following network:
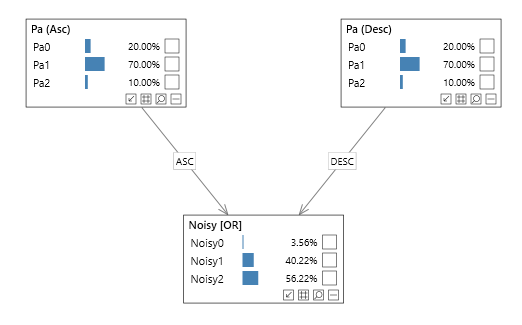
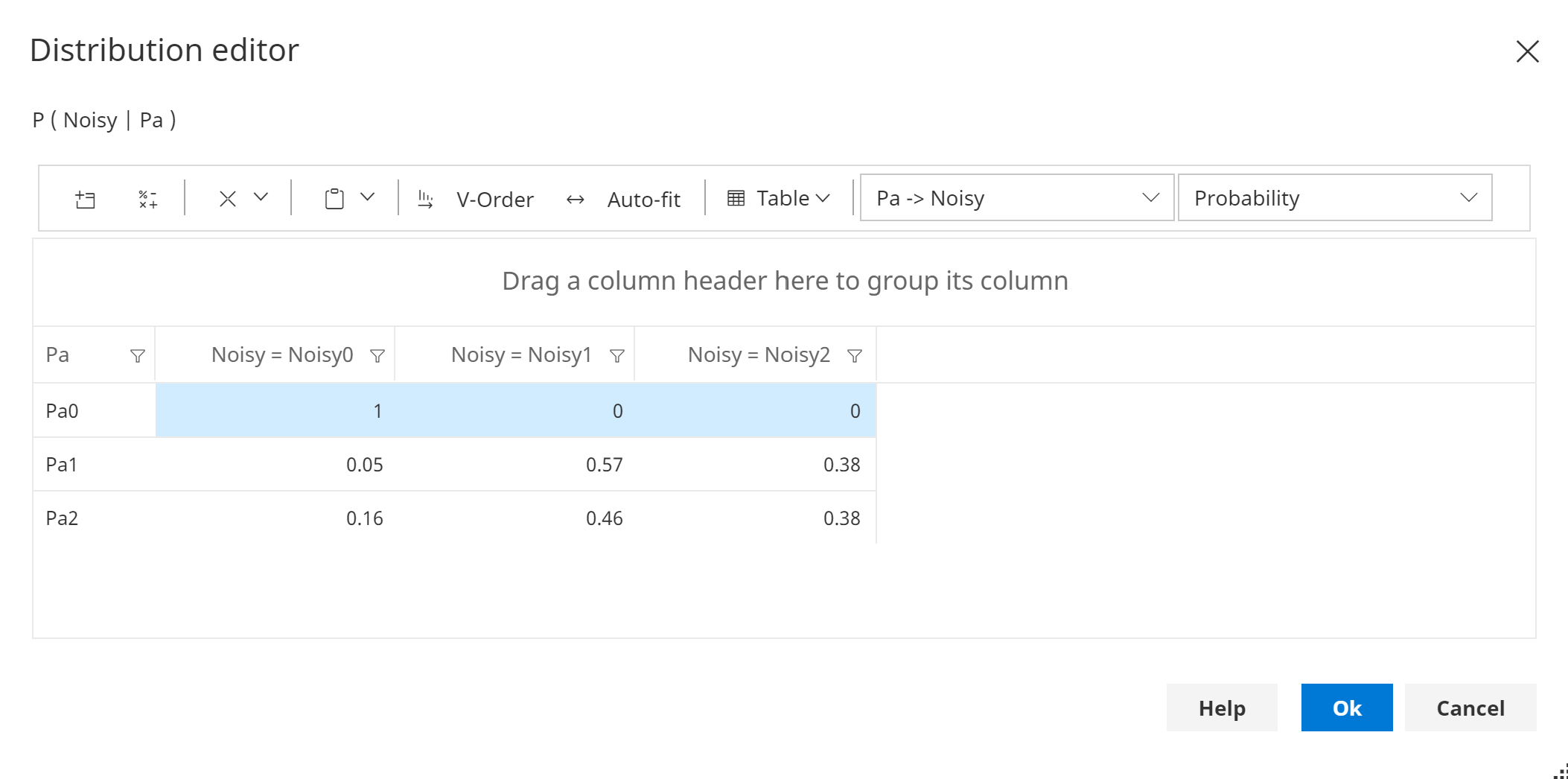
Noisy order - Ascending
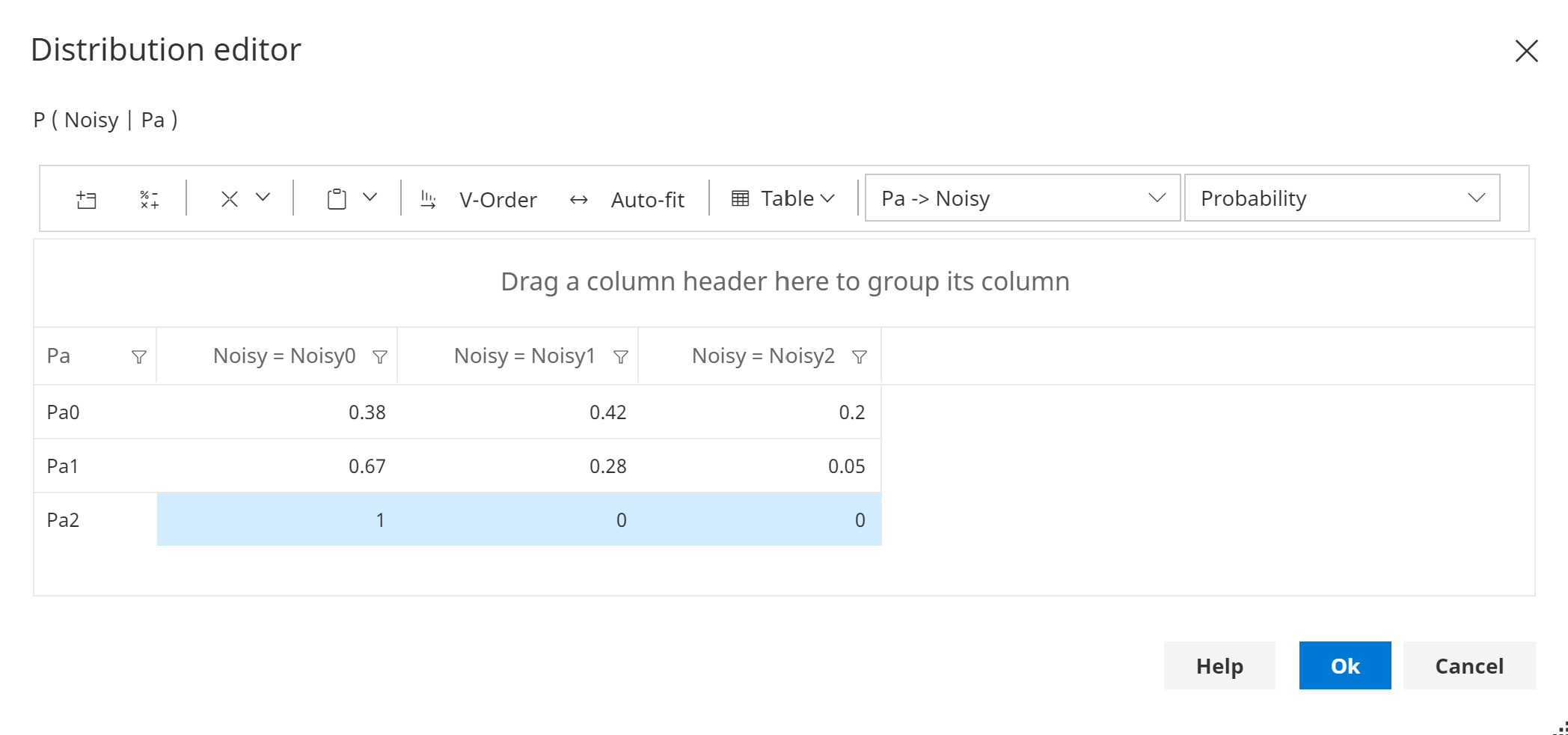
The first state of Pa is the 'no influence' (designated) state for Noisy, as shown in the image below.
The highlighted cells are constrained values, and hence cannot be changed.
Noisy order - Descending
The last state of Pa is the 'no influence' (designated) state for Noisy, as shown in the image below.
The highlighted cells are constrained values, and cannot be changed.
The reason both Ascending and Descending are supported, rather than simply imposing an order, is that a parent Pa may be parent of multiple noisy nodes, and influence them in different orders.
The Noisy order can be changed by clicking on a link, and setting the order in the Properties pane.
Leak
An example of the leak distribution is shown below:
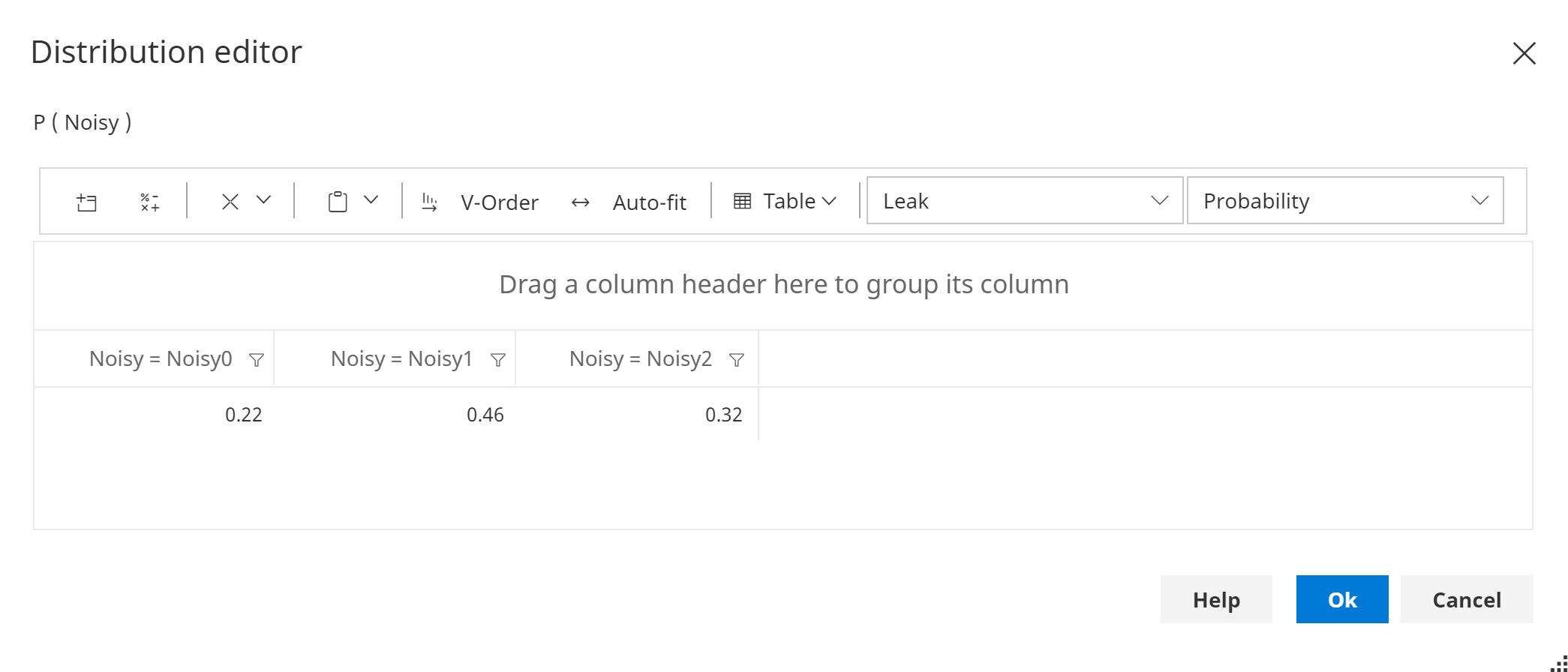
If there are no other factors to consider other than the existing parents of the noisy node, this distribution can be set to {1, 0, 0, ..., 0}.
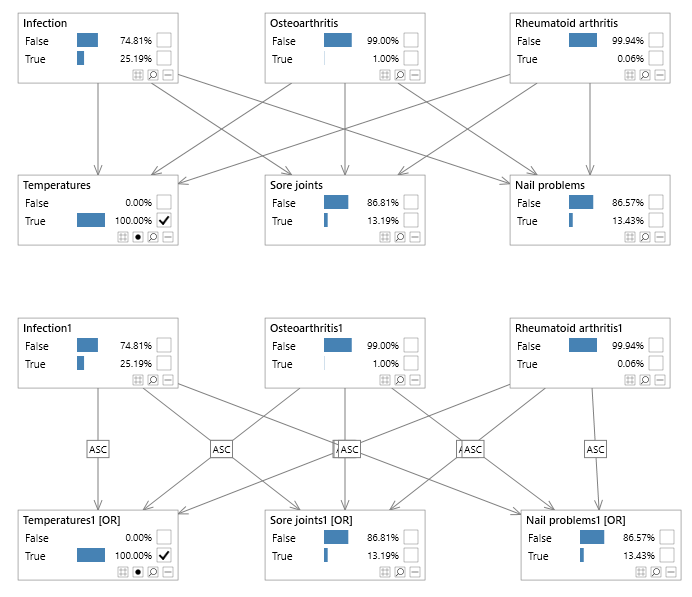
Example
Consider the two Bayesian networks in the image below (see the NoisyOr network included with Bayes Server, available from the Start page). The bottom network contains Noisy nodes whereas the top one does not. The top network has 27 parameters but requires all disease combinations to be defined for each symptom, whereas the noisy network has only 15 parameters which makes it less expressive but a more scalable architecture when the assumptions of causal independence are valid or 'good enough'.